|
Disclaimer: This work has been submitted by a student. This is not an example of the work produced by our Essay Writing Service. You can view samples of our professional work here.
How To Do Gram Staining. The Gram-staining process discovered in 1882 (published 1884) by Hans Christian Gram, a Danish bacteriologist and plays an important role in the classifying the bacteria. Gram-staining is usually the first step in identification bacteria and can be used in characterizing bacteria. Escherichia coli. Escherichia coli (commonly abbreviated E. Coli) is a Gram-negative, rod-shaped bacterium that is commonly found in the lower intestine of warm-blooded organisms (endotherms). Coli strains are harmless, but some serotypes can cause serious food poisoning in humans.
Any opinions, findings, conclusions or recommendations expressed in this material are those of the authors and do not necessarily reflect the views of UK Essays.
Observation of microorganism under microscope can be improved by using certain processes and techniques such as the staining. Staining is an important step to observe microorganisms more clearly, to differentiate between microorganisms as well as to differentiate parts in microorganism (Bagyaraj et al, 2005). The identification, morphology, some extracellular and intracellular components of microorganisms can be determined and detected through the staining. Many microorganisms difficult to be observed under microscope due to their colourless appearance and semitransparent properties as their refractive index almost same as surroundings (Patil et al, 2008). The stain improves contrast for visualizing microorganisms. Staining process can be explained either as physical, chemical reaction or combination of the both reaction.
There are different types of staining such as the simple stain, differential stain and special stain. Simple stain can be used for observing certain basic structures as well as the shape of microorganisms. Differential stain while can be used in distinguishing between different types of microorganisms. Special stain on the other hand can be used for identifying specific structures in the microorganisms such as the flagella (Frey & Price, 2003). Gram-stain is one of the commonly used differential stains. The Gram-staining process discovered in 1882 (published 1884) by Hans Christian Gram, a Danish bacteriologist and plays an important role in the classifying the bacteria. Gram-staining is usually the first step in identification bacteria and can be used in characterizing bacteria. Bacteria species can be separated into two large groups, which are the Gram-positive and Gram-negative groups through the Gram-staining (Sridhar Rao, n.d.). This process also important in clinical laboratory such as to examine and identify bacteria responsible for certain diseases.
Staining process requires the preparation of smear that contains a thin layer of bacteria. The preparation of smear involves spreading and fixing of microorganisms on the microscope slide. Use of smear prevents microorganisms from being washing away with stain (Vasanthakumari, 2009). Besides the smear, there are four important components in the Gram stain process, which are the primary stain, mordant, decolourizing agent as well as the counterstain that used in sequences. The primary stains usually basic dye such as crystal violet that reacts with acidic component of cell and causes all the bacteria to be stained with the crystal violet or purple. The other dye like the methyl violet can also be used. The other component, mordant in the Gram stain refers to iodine. Mordant is chemical that increases affinity of the stain to the microorganisms and also their coating, making certain structures thicker for easier observation under microscope. The decolorizing agent decolorizes dye from cell that already being stained (Rajan, 2005). The degree of decolorization different in bacteria depends on their chemical components. Decolourization agent commonly refers to ethanol or other solution like acetone or mixture of acetone and ethyl alcohol. Counterstain while is another basic dye that important in giving new colour for cells that decolourized. Counterstain can be the safranin (used in this practical) or the carbon fuchsin.
The Gram stain (differential stains) gives different colour for different types of bacteria. The colour is the one that determine whether the bacterium is Gram positive or Gram negative. The Gram positive bacteria resist decolourization and give result of crystal violet or purple colour (primary stain). Gram-negative bacteria decolorize and give red or pink colour as it takes up counterstain (Ananthanarayan & Paniker, 2006). The difference in result is due to the differences in the cell wall structure or composition of bacteria that causes the different in the reaction with the series of reagents in Gram staining (Talaro, 2007).
Get help with your essay today, from our professional essay writers!
Qualified writers in the subject of biology are ready and waiting to help you with your studies.
Get help with your essay today
from our professional essay writers!
Visit www.ukessays.com to see how we can help you!Preparation of Staining Reagents:
Crystal violet
Solution A: Crystal violet 2.0g
Ethanol, 95% (v/v) 20 ml
Solution B: Ammonium oxalate 0.8g
Distilled water 80 ml
Solution A and B mixed.
Mordant
Iodine 1.0 g
Potassium iodide 2.0 g
Distilled water 300 ml
Iodine and potassium blended with mortar, distilled water added during blending until iodine dissolved.
Decolorization solvent
Ethanol, 95% (v/v)
Counterstain
Safranin 0.25 g [2.5 %(w/v)]
Ethanol 10 ml [9.5% (v/v)]
Distilled water 90 ml
Materials:
Glass slide
Escherichia coli in broth culture
Escherichia coli in agar culture
Bacillus sp. in broth culture
Bacillus sp. in agar culture
Staphylococcus aureus in broth culture
Actinomycetes sp. in broth culture
Actinomycetes sp. in agar culture
Kimwipe
Bunsen burner
Dropper
Distilled water
Inoculation loop
Procedure:
Preparation of smear:
For culture taken from liquid medium (broth), 1 drop of culture to be examined was transferred by using inoculation loop onto a slide and spread to from circular smear. For culture taken from solid medium (agar), one drop of distilled water first dispensed on the slide. The single colony then spread on the water to form circular smear.
The slide was heat-fixed with flame.
Gram-staining
The slide was placed on the rack.
1-2 drops of crystal violet was dropped on the smear and left for 2 minutes.
The crystal violet was rinsed off with distilled water for 2 seconds.
Iodine solution was dropped and left for 2 minutes.
The iodine solution was rinsed off with distilled water for 2 seconds.
The smear was decolorized by washing with ethanol (95%v/v) for less than 10 seconds. The ethanol then rinsed off with distilled water for 10 seconds.
Safranin solution was dropped on the smear for 10 seconds.
The red-coloured safranin was rinsed-off with distilled water.
The side was dried using Kimwipe or air-dry.
The slide was observed under the microscope.
Results:(A)Escherichia coli
G:DCIM101NIKONDSCN1773.JPG
1(a) Broth culture (zoom in).
1(b) Agar plate (zoom in).
Figure 1: Microscopic image of Escherichia coli under total magnification of 400Ã- from different culture
(B) Bacillus species
G:DCIM101NIKONDSCN1745.JPG G:DCIM101NIKONDSCN1738.JPG
2(a) Broth culture (zoom in).
2(b) Agar plate (zoom in).
Figure 2: Microscopic image of Bacillus sp. under total magnification of 400Ã- from different cultures.
(C) Staphylococcus Aureus
G:DCIM101NIKONDSCN1767.JPG
Figure 3: Microscopic image of Staphylococcus aureus under total magnification of 400Ã- from broth culture (zoom in).
(D) Actinomycetes species
C:UsersmichelleDocumentsUMS MICROBIOLOGYPHOTOSS1.JPG G:DCIM101NIKONDSCN1760.JPG
4(a) Broth culture (zoom in) under total magnification of 400Ã-.
4(b) Agar plate (zoom in) under total magnification of 400Ã-.
Figure 3: Microscopic image of Actinomycetes sp. under different magnification from different culture.
Table 1: The result of Gram stain on different microorganism
Type of microorganismsShape of the microorganismsColour stained on microorganismsGram positive or Gram negative
Escherichia coli (broth culture)
Bacillus or Rod-shaped
Pink
Gram negative
Escherichia coli (agar plate)
Bacillus or Rod-shaped
Pink
Gram negative
Bacillus sp. (broth culture)
Bacillus or Rod-shaped
Purple
Gram positive
Bacillus sp. (agar plate)
Bacillus or Rod-shaped
Purple
Gram positive
Staphylococcus aureus
Coccus or round-shaped
Purple
Gram positive
Actinomycetes sp. (broth culture)
Mycelial
Purple
Gram positive
Actinomycetes sp. (agar plate)
Mycelial
Purple
Gram positive
Discussion:
For every bacterium studied, a smear is first prepared as the smear enables Gram staining to be done without washing away bacteria together with stain. The spreading process (for both broth and agar culture) enables the distribution of bacteria on slides so that suitable density of bacteria can be found on the slide. This increases chance of individual bacteria to be observed under microscope (Port, 2009). The microorganisms from agar first suspended in distilled water before spreading. Without spreading, bacteria may be too concentrated, crowded and overlapped (in clumps), making the observation to be difficult. The slide was heat fixed after drying. Heating enables coagulation and precipitation of protein of bacteria to occurs, hence fix the bacteria on slide. The bacteria killed and adhere to the surface. Fixation makes the bacteria rigid, immobile, increased permeability and affinity to staining. This also prevents the autolysis process of bacteria (Aneja, 2003). During the fixation process, slides not be placed directly above the heat or passed through too many times as overheat may causes changes in the shape and hence cause the distortion of the microorganisms. At the same time, less heat supplied may cause the microorganisms do not fix firmly. Before heat fix, the slide is allowed to dry completely as wet bacterial suspension may create aerosol (Shimeld, 1999).The presence of water may also cause over heating.
Get help with your essay today, from our professional essay writers!
Qualified writers in the subject of biology are ready and waiting to help you with your studies.
Get help with your essay today
from our professional essay writers!
Visit www.ukessays.com to see how we can help you!
The crystal violet added as the primary stain. Crystal violet is basic dye and has affinity for cell structures that are acidic such as the protoplasm. Crystal violet is added to stain everything on slide or to stain all bacteria (Gram positive or Gram negative). This is same for all the seven samples. Crystal violet dye enters the cells and stained with crystal violet colour. It was suggested that the aqueous dye dissociated into CV+ ion and chloride, Cl- ion (Hussey & Smith, n.d.). The positively charged ion binds to the negatively charged components in cell after penetrating the cell wall and cell membrane, hence giving the purple colour. The extra crystal violet dye that not binds to cell is cleared by distilled water. Addition of iodine in next step enables the crystal violet dye to further fix and adhere to organisms (Medical Education Division, 2006). This is due to the formation of complex between iodine and dye ion (CV-I complex) as the negatively charged iodine ion (I- or I3- ion) binds to the positively charged ion of dye (CV+ ion) in cytoplasm and hence bacteria appeared as violet colour (Vasanthakumari, 2009). The solubility of the dye decreased during the process as the ions bind to organisms. Iodine acts as mordant as it increases affinity of crystal violet stain to organisms.
The addition of 95% ethanol as decolourizer enables the lipid to be extracted or dissolved from the cell wall for the Gram negative bacteria like the Escherichia coli. Gram negative bacteria have an outer membrane that constitutes most of the cell wall, also known as lipopolysaccharide layer (LPS) in cell wall (Clark et al, 2009). This is a lipid bilayer structure that differs from cytoplasmic membrane. This layer not only made up of phospholipids and protein, but also polysaccharides that not commonly found in cytoplasmic membrane. Polysaccharide portion made up of core polysaccharides and O-polysaccharides while the lipid portion made up of lipid A which then bind to the core polysaccharides. This LPS layer is located outside a thin layer of peptidoglycan. The outer membrane gives rises to high lipid composition in the cell wall. Decolourizer dissolve off lipid, hence increases the permeability of cell wall which eventually enables the crystal violet-iodine complex to be lost together with the lipid.
The cell wall (murein layer) of Gram positive layer while has no outer membrane but have thick, cross-linked and multi-layered peptidoglycan. Teichoic acids, the phosphorylated polyalcohol can be found embedded in peptidoglycan layers. These acids can be found bonded to muramic acid residues in peptidoglycan. Lipoteichoic acid which refers to the teichoic acids that binds to the lipids of membrane can also be found in Gram positive bacterial cell wall. In certain actinobacteria, structure called mycolic acids also can be found. The lack of outer membrane gives rises to low lipid composition in cell wall. Hence, the action of decolorizer on Gram positive bacteria (Bacillus sp., Staphylococcus aureus and Actinomycetes sp.) causes dehydration of cell wall due to the thick peptidoglycan and the composition of lipid available to be dissolved is low. This eventually decreases cell wall permeability, closing pores on cell wall and hence retain the crystal violet-iodine complex inside (Differential staining: The Gram Stain, n.d.). As the cell shrinks, the complex trapped in the thick peptidoglycan and hence cells do not decolourized. After this process, E. coli is in colourless as the crystal-violet iodine complex loses while Bacillus sp., Staphylococcus aureus and Actinomycetes sp. still in purple colour.
Ethanol was not added for more than 30 seconds. Over decolourization can cause the stain of Gram positive bacteria to decolourize and appears as Gram negative (Betts et al, 2003). Under decolourization (too short) also avoided as it can cause dye to be removed incompletely from Gram negative bacteria. Both situations can give false results. After decolorization, smear was washed with distilled water for 15 second to completely stop the decolourization process. The counterstain, safranin solution then stained the E. coli that is colourless with the red colour. Safranin is basic dye (cationic ion) carry the positive dye ion, chromophore that attached to acidic cell structures (negatively charged) such as the protoplasm. Basic dye also attached to other negatively charged macromolecules like proteins and nucleic acid (Archunan, 2004). Both the Gram positive and Gram negative bacteria took up the counterstain but the colour of Gram Positive do not change much as it already stained with purple. For every dye, there is different period of time for staining. This is to prevent over or under stain that may results in inaccurate result.
From the observation, Escherichia coli stained red and give accurate result of Gram negative. The shape of E. coli can be observed as rod shape. Bacillus sp., Staphylococcus aureus and Actinomycetes sp. while shows results of Gram positive as all are stained with purple colour. The shapes observed are respectively rod-shaped, round-shaped and in mycelial. For Staphylococcus aureus, the cocci shape is sticked together in clumps or amorphous sheet and not separated. For E. coli, bacillus sp. and staphylococcus aureus, two samples are taken, one from the broth and one from the agar. Both the samples show the same results. The difference is on the amount of microorganisms observed. Bacillus sp., for example, that taken from agar plate is very crowded. This is because the each colony taken contains a number of microorganisms. It is more difficult to be observed the shape of the organisms. However, the colour stained can be observed clearly. For the broth culture, individual organisms and the shape as well as the colour can be observed more clearly.
Get help with your essay today, from our professional essay writers!
Qualified writers in the subject of biology are ready and waiting to help you with your studies.
Get help with your essay today
from our professional essay writers!
Visit www.ukessays.com to see how we can help you!Conclusion:
Gram staining is important in differentiating Gram positive and Gram negative bacteria in which the Gram positive bacteria stained purple colour while Gram negative organisms stained pink. Escherichia coli is Gram negative while bacillus sp., staphylococcus aureus and actinomycetes are Gram positive bacteria.
Escherichia coli (/ˌɛʃəˈrɪkiəˈkoÊŠlaɪ/),[1][2] also known as E. coli (/ËŒiËˈkoÊŠlaɪ/),[3] is a Gram-negative, facultative anaerobic, rod-shaped, coliform bacterium of the genusEscherichia that is commonly found in the lower intestine of warm-blooded organisms (endotherms).[4][5] Most E. colistrains are harmless, but some serotypes can cause serious food poisoning in their hosts, and are occasionally responsible for product recalls due to food contamination.[6][7] The harmless strains are part of the normal microbiota of the gut, and can benefit their hosts by producing vitamin K2,[8] and preventing colonization of the intestine with pathogenic bacteria, having a symbiotic relationship.[9][10]E. coli is expelled into the environment within fecal matter. The bacterium grows massively in fresh fecal matter under aerobic conditions for 3 days, but its numbers decline slowly afterwards.[11]
E. coli and other facultative anaerobes constitute about 0.1% of gut microbiota,[12] and fecal–oral transmission is the major route through which pathogenic strains of the bacterium cause disease. Cells are able to survive outside the body for a limited amount of time, which makes them potential indicator organisms to test environmental samples for fecal contamination.[13][14] A growing body of research, though, has examined environmentally persistent E. coli which can survive for extended periods outside a host.[15]
The bacterium can be grown and cultured easily and inexpensively in a laboratory setting, and has been intensively investigated for over 60 years. E. coli is a chemoheterotroph whose chemically defined medium must include a source of carbon and energy.[16]E. coli is the most widely studied prokaryoticmodel organism, and an important species in the fields of biotechnology and microbiology, where it has served as the host organism for the majority of work with recombinant DNA. Under favorable conditions, it takes up to 20 minutes to reproduce.[17]
Biology and biochemistry[edit]
Model of successive binary fission in E. coli
A colony of E. coli growing
Type and morphology[edit]
E. coli is a Gram-negative, facultative anaerobe (that makes ATP by aerobic respiration if oxygen is present, but is capable of switching to fermentation or anaerobic respiration if oxygen is absent) and nonsporulating bacterium.[18] Cells are typically rod-shaped, and are about 2.0 μm long and 0.25–1.0 μm in diameter, with a cell volume of 0.6–0.7 μm3.[19][20][21]
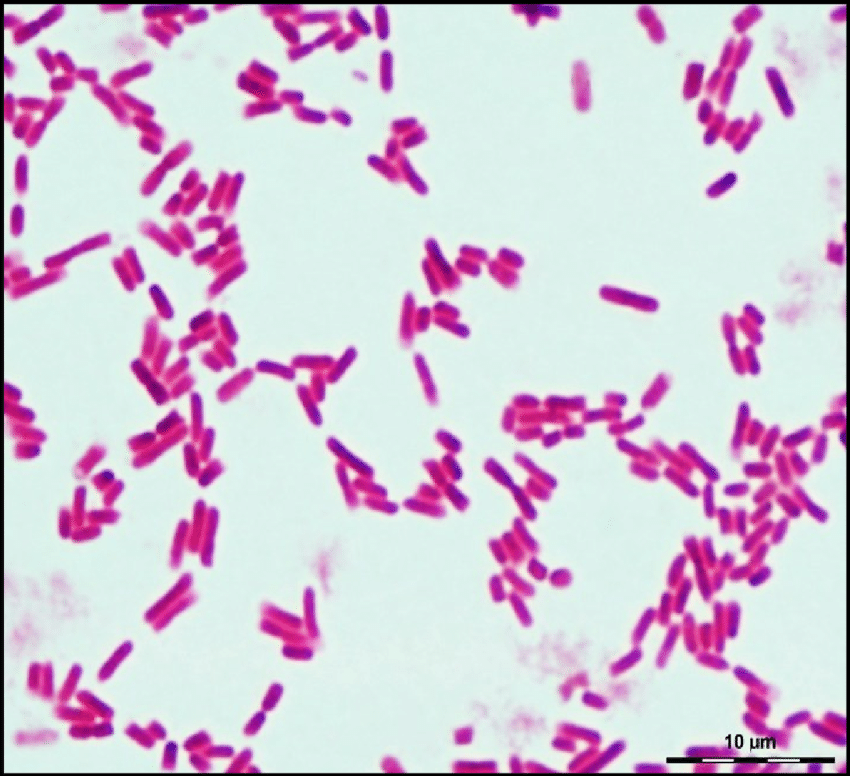
E. coli stains Gram-negative because its cell wall is composed of a thin peptidoglycan layer and an outer membrane. During the staining process, E. coli picks up the color of the counterstain safranin and stains pink. The outer membrane surrounding the cell wall provides a barrier to certain antibiotics such that E. coli is not damaged by penicillin.[16]
Strains that possess flagella are motile. The flagella have a peritrichous arrangement.[22] It also attaches and effaces to the microvilli of the intestines via an adhesion molecule known as intimin.[23]
Metabolism[edit]
E. coli can live on a wide variety of substrates and uses mixed-acid fermentation in anaerobic conditions, producing lactate, succinate, ethanol, acetate, and carbon dioxide. Since many pathways in mixed-acid fermentation produce hydrogen gas, these pathways require the levels of hydrogen to be low, as is the case when E. coli lives together with hydrogen-consuming organisms, such as methanogens or sulphate-reducing bacteria.[24]
Culture growth[edit]
Optimum growth of E. coli occurs at 37 °C (98.6 °F), but some laboratory strains can multiply at temperatures up to 49 °C (120 °F).[25]E. coli grows in a variety of defined laboratory media, such as lysogeny broth, or any medium that contains glucose, ammonium phosphate monobasic, sodium chloride, magnesium sulfate, potassium phosphate dibasic, and water. Growth can be driven by aerobic or anaerobic respiration, using a large variety of redox pairs, including the oxidation of pyruvic acid, formic acid, hydrogen, and amino acids, and the reduction of substrates such as oxygen, nitrate, fumarate, dimethyl sulfoxide, and trimethylamine N-oxide.[26]E. coli is classified as a facultative anaerobe. It uses oxygen when it is present and available. It can, however, continue to grow in the absence of oxygen using fermentation or anaerobic respiration. The ability to continue growing in the absence of oxygen is an advantage to bacteria because their survival is increased in environments where water predominates.[16]
Cell cycle[edit]
The bacterial cell cycle is divided into three stages. The B period occurs between the completion of cell division and the beginning of DNA replication. The C period encompasses the time it takes to replicate the chromosomal DNA. The D period refers to the stage between the conclusion of DNA replication and the end of cell division.[27] The doubling rate of E. coli is higher when more nutrients are available. However, the length of the C and D periods do not change, even when the doubling time becomes less than the sum of the C and D periods. At the fastest growth rates, replication begins before the previous round of replication has completed, resulting in multiple replication forks along the DNA and overlapping cell cycles.[28]
Genetic adaptation[edit]
E. coli and related bacteria possess the ability to transfer DNA via bacterial conjugation or transduction, which allows genetic material to spread horizontally through an existing population. The process of transduction, which uses the bacterial virus called a bacteriophage,[29] is where the spread of the gene encoding for the Shiga toxin from the Shigella bacteria to E. coli helped produce E. coli O157:H7, the Shiga toxin-producing strain of E. coli.
Diversity[edit]
Scanning electron micrograph of an E. coli colony.
E. coli encompasses an enormous population of bacteria that exhibit a very high degree of both genetic and phenotypic diversity. Genome sequencing of a large number of isolates of E. coli and related bacteria shows that a taxonomic reclassification would be desirable. However, this has not been done, largely due to its medical importance,[30] and E. coli remains one of the most diverse bacterial species: only 20% of the genes in a typical E. coli genome is shared among all strains.[31]
In fact, from the evolutionary point of view, the members of genus Shigella (S. dysenteriae, S. flexneri, S. boydii, and S. sonnei) should be classified as E. coli strains, a phenomenon termed taxa in disguise.[32] Similarly, other strains of E. coli (e.g. the K-12 strain commonly used in recombinant DNA work) are sufficiently different that they would merit reclassification.
A strain is a subgroup within the species that has unique characteristics that distinguish it from other strains. These differences are often detectable only at the molecular level; however, they may result in changes to the physiology or lifecycle of the bacterium. For example, a strain may gain pathogenic capacity, the ability to use a unique carbon source, the ability to take upon a particular ecological niche, or the ability to resist antimicrobial agents. Different strains of E. coli are often host-specific, making it possible to determine the source of fecal contamination in environmental samples.[13][14] For example, knowing which E. coli strains are present in a water sample allows researchers to make assumptions about whether the contamination originated from a human, another mammal, or a bird.
Serotypes[edit]
A common subdivision system of E. coli, but not based on evolutionary relatedness, is by serotype, which is based on major surface antigens (O antigen: part of lipopolysaccharide layer; H: flagellin; K antigen: capsule), e.g. O157:H7).[33] It is, however, common to cite only the serogroup, i.e. the O-antigen. At present, about 190 serogroups are known.[34] The common laboratory strain has a mutation that prevents the formation of an O-antigen and is thus not typeable.
Genome plasticity and evolution[edit]
Like all lifeforms, new strains of E. colievolve through the natural biological processes of mutation, gene duplication, and horizontal gene transfer; in particular, 18% of the genome of the laboratory strain MG1655 was horizontally acquired since the divergence from Salmonella.[35]E. coli K-12 and E. coli B strains are the most frequently used varieties for laboratory purposes. Some strains develop traits that can be harmful to a host animal. These virulent strains typically cause a bout of diarrhea that is often self-limiting in healthy adults but is frequently lethal to children in the developing world.[36] More virulent strains, such as O157:H7, cause serious illness or death in the elderly, the very young, or the immunocompromised.[36][37]
The genera Escherichia and Salmonella diverged around 102 million years ago (credibility interval: 57–176 mya), which coincides with the divergence of their hosts: the former being found in mammals and the latter in birds and reptiles.[38] This was followed by a split of an Escherichia ancestor into five species (E. albertii, E. coli, E. fergusonii, E. hermannii, and E. vulneris). The last E. coli ancestor split between 20 and 30 million years ago.[39]
The long-term evolution experiments using E. coli, begun by Richard Lenski in 1988, have allowed direct observation of genome evolution over more than 65,000 generations in the laboratory.[40] For instance, E. coli typically do not have the ability to grow aerobically with citrate as a carbon source, which is used as a diagnostic criterion with which to differentiate E. coli from other, closely, related bacteria such as Salmonella. In this experiment, one population of E. coli unexpectedly evolved the ability to aerobically metabolize citrate, a major evolutionary shift with some hallmarks of microbial speciation.
Neotype strain[edit]
E. coli is the type species of the genus (Escherichia) and in turn Escherichia is the type genus of the family Enterobacteriaceae, where the family name does not stem from the genus Enterobacter + 'i' (sic.) + 'aceae', but from 'enterobacterium' + 'aceae' (enterobacterium being not a genus, but an alternative trivial name to enteric bacterium).[41][42][43]
The original strain described by Escherich is believed to be lost, consequently a new type strain (neotype) was chosen as a representative: the neotype strain is U5/41T,[44] also known under the deposit names DSM 30083,[45]ATCC 11775,[46] and NCTC 9001,[47] which is pathogenic to chickens and has an O1:K1:H7 serotype.[48] However, in most studies, either O157:H7, K-12 MG1655, or K-12 W3110 were used as a representative E. coli. The genome of the type strain has only lately been sequenced.[44]
Phylogeny of E. coli strains[edit]
A large number of strains belonging to this species have been isolated and characterised. In addition to serotype (vide supra), they can be classified according to their phylogeny, i.e. the inferred evolutionary history, as shown below where the species is divided into six groups.[49][50] Particularly the use of whole genome sequences yields highly supported phylogenies. Based on such data, five subspecies of E. coli were distinguished.[44]
The link between phylogenetic distance ('relatedness') and pathology is small,[44]e.g. the O157:H7 serotype strains, which form a clade ('an exclusive group')—group E below—are all enterohaemorragic strains (EHEC), but not all EHEC strains are closely related. In fact, four different species of Shigella are nested among E. coli strains (vide supra), while E. albertii and E. fergusonii are outside this group.Indeed, all Shigella species were placed within a single subspecies of E. coli in a phylogenomic study that included the type strain,[44] and for this reason an according reclassification is difficult.All commonly used research strains of E. coli belong to group A and are derived mainly from Clifton's K-12 strain (λ⺠Fâº; O16) and to a lesser degree from d'Herelle's Bacillus coli strain (B strain)(O7).
Genomics[edit]
An image of E.coli using early electron microscopy.
The first complete DNA sequence of an E. coligenome (laboratory strain K-12 derivative MG1655) was published in 1997. It is a circular DNA molecule 4.6 million base pairs in length, containing 4288 annotated protein-coding genes (organized into 2584 operons), seven ribosomal RNA (rRNA) operons, and 86 transfer RNA (tRNA) genes. Despite having been the subject of intensive genetic analysis for about 40 years, a large number of these genes were previously unknown. The coding density was found to be very high, with a mean distance between genes of only 118 base pairs. The genome was observed to contain a significant number of transposable genetic elements, repeat elements, cryptic prophages, and bacteriophage remnants.[51]
More than three hundred complete genomic sequences of Escherichia and Shigella species are known. The genome sequence of the type strain of E. coli was added to this collection before 2014.[44] Comparison of these sequences shows a remarkable amount of diversity; only about 20% of each genome represents sequences present in every one of the isolates, while around 80% of each genome can vary among isolates.[31] Each individual genome contains between 4,000 and 5,500 genes, but the total number of different genes among all of the sequenced E. coli strains (the pangenome) exceeds 16,000. This very large variety of component genes has been interpreted to mean that two-thirds of the E. colipangenome originated in other species and arrived through the process of horizontal gene transfer.[52]
Gene nomenclature[edit]
Genes in E. coli are usually named by 4-letter acronyms that derive from their function (when known) and italicized. For instance, recA is named after its role in homologous recombination plus the letter A. Functionally related genes are named recB, recC, recD etc. The proteins are named by uppercase acronyms, e.g. RecA, RecB, etc. When the genome of E. coli was sequenced, all genes were numbered (more or less) in their order on the genome and abbreviated by b numbers, such as b2819 (= recD). The 'b' names were created after Fred Blattner, who led the genome sequence effort.[53] Another numbering system was introduced with the sequence of another E. coli strain, W3110, which was sequenced in Japan and hence uses numbers starting by JW... (Japanese W3110), e.g. JW2787 (= recD).[54] Hence, recD = b2819 = JW2787. Note, however, that most databases have their own numbering system, e.g. the EcoGene database[55] uses EG10826 for recD. Finally, ECK numbers are specifically used for alleles in the MG1655 strain of E. coli K-12.[55] Complete lists of genes and their synonyms can be obtained from databases such as EcoGene or Uniprot.
Proteomics[edit]Proteome[edit]
Several studies have investigated the proteome of E. coli. By 2006, 1,627 (38%) of the 4,237 open reading frames (ORFs) had been identified experimentally.[56]The 4,639,221–base pair sequence of Escherichia coli K-12 is presented. Of 4288 protein-coding genes annotated, 38 percent have no attributed function. Comparison with five other sequenced microbes reveals ubiquitous as well as narrowly distributed gene families; many families of similar genes within E. coli are also evident. The largest family of paralogous proteins contains 80 ABC transporters. The genome as a whole is strikingly organized with respect to the local direction of replication; guanines, oligonucleotides possibly related to replication and recombination, and most genes are so oriented. The genome also contains insertion sequence (IS) elements, phage remnants, and many other patches of unusual composition indicating genome plasticity through horizontal transfer.[57]
Interactome[edit]
The interactome of E. coli has been studied by affinity purification and mass spectrometry (AP/MS) and by analyzing the binary interactions among its proteins.
Protein complexes. A 2006 study purified 4,339 proteins from cultures of strain K-12 and found interacting partners for 2,667 proteins, many of which had unknown functions at the time.[58] A 2009 study found 5,993 interactions between proteins of the same E. coli strain, though these data showed little overlap with those of the 2006 publication.[59]
Binary interactions. Rajagopala et al. (2014) have carried out systematic yeast two-hybrid screens with most E. coli proteins, and found a total of 2,234 protein-protein interactions.[60] This study also integrated genetic interactions and protein structures and mapped 458 interactions within 227 protein complexes.
Normal microbiota[edit]
E. coli belongs to a group of bacteria informally known as coliforms that are found in the gastrointestinal tract of warm-blooded animals.[41]E. coli normally colonizes an infant's gastrointestinal tract within 40 hours of birth, arriving with food or water or from the individuals handling the child. In the bowel, E. coli adheres to the mucus of the large intestine. It is the primary facultative anaerobe of the human gastrointestinal tract.[61] (Facultative anaerobes are organisms that can grow in either the presence or absence of oxygen.) As long as these bacteria do not acquire genetic elements encoding for virulence factors, they remain benign commensals.[62]
Therapeutic use[edit]
Nonpathogenic E. coli strain Nissle 1917, (Mutaflor) and E. coli O83:K24:H31 (Colinfant)[63][64]) are used as probiotic agents in medicine, mainly for the treatment of various gastrointestinal diseases,[65] including inflammatory bowel disease.[66]
Role in disease[edit]
Most E. coli strains do not cause disease, naturally living in the gut,[67] but virulent strains can cause gastroenteritis, urinary tract infections, neonatalmeningitis, hemorrhagic colitis, and Crohn's disease. Common signs and symptoms include severe abdominal cramps, diarrhea, hemorrhagic colitis, vomiting, and sometimes fever. In rarer cases, virulent strains are also responsible for bowel necrosis (tissue death) and perforation without progressing to hemolytic-uremic syndrome, peritonitis, mastitis, sepsis, and Gram-negative pneumonia. Very young children are more susceptible to develop severe illness, such as hemolytic uremic syndrome; however, healthy individuals of all ages are at risk to the severe consequences that may arise as a result of being infected with E. coli.[61][68][69][70]
Some strains of E. coli, for example O157:H7, can produce Shiga toxin (classified as a bioterrorism agent). The Shiga toxin causes inflammatory responses in target cells of the gut, leaving behind lesions which result in the bloody diarrhea that is a symptom of a Shiga toxin-producing E. coli (STEC) infection. This toxin further causes premature destruction of the red blood cells, which then clog the body's filtering system, the kidneys, in some rare cases (usually in children and the elderly) causing hemolytic-uremic syndrome (HUS), which may lead to kidney failure and even death. Signs of hemolytic uremic syndrome include decreased frequency of urination, lethargy, and paleness of cheeks and inside the lower eyelids. In 25% of HUS patients, complications of nervous system occur, which in turn causes strokes. In addition, this strain causes the buildup of fluid (since the kidneys do not work), leading to edema around the lungs and legs and arms. This increase in fluid buildup especially around the lungs impedes the functioning of the heart, causing an increase in blood pressure.[71][23][72][73][74][69][70]
Uropathogenic E. coli (UPEC) is one of the main causes of urinary tract infections.[75] It is part of the normal microbiota in the gut and can be introduced in many ways. In particular for females, the direction of wiping after defecation (wiping back to front) can lead to fecal contamination of the urogenital orifices. Anal intercourse can also introduce this bacterium into the male urethra, and in switching from anal to vaginal intercourse, the male can also introduce UPEC to the female urogenital system.
Enterotoxigenic E. coli (ETEC) is the most common cause of traveler's diarrhea, with as many as 840 million cases worldwide in developing countries each year. The bacteria, typically transmitted through contaminated food or drinking water, adheres to the intestinal lining, where it secretes either of two types of enterotoxins, leading to watery diarrhea. The rate and severity of infections are higher among children under the age of five, including as many as 380,000 deaths annually.[76]
In May 2011, one E. coli strain, O104:H4, was the subject of a bacterial outbreak that began in Germany. Certain strains of E. coli are a major cause of foodborne illness. The outbreak started when several people in Germany were infected with enterohemorrhagic E. coli (EHEC) bacteria, leading to hemolytic-uremic syndrome (HUS), a medical emergency that requires urgent treatment. The outbreak did not only concern Germany, but also 15 other countries, including regions in North America.[77] On 30 June 2011, the German Bundesinstitut für Risikobewertung (BfR) (Federal Institute for Risk Assessment, a federal institute within the German Federal Ministry of Food, Agriculture and Consumer Protection) announced that seeds of fenugreek from Egypt were likely the cause of the EHEC outbreak.[78]
Incubation period[edit]
The time between ingesting the STEC bacteria and feeling sick is called the 'incubation period'. The incubation period is usually 3–4 days after the exposure, but may be as short as 1 day or as long as 10 days. The symptoms often begin slowly with mild belly pain or non-bloody diarrhea that worsens over several days. HUS, if it occurs, develops an average 7 days after the first symptoms, when the diarrhea is improving.[79]
Treatment[edit]
The mainstay of treatment is the assessment of dehydration and replacement of fluid and electrolytes. Administration of antibiotics has been shown to shorten the course of illness and duration of excretion of enterotoxigenic E. coli (ETEC) in adults in endemic areas and in traveller's diarrhea, though the rate of resistance to commonly used antibiotics is increasing and they are generally not recommended.[80] The antibiotic used depends upon susceptibility patterns in the particular geographical region. Currently, the antibiotics of choice are fluoroquinolones or azithromycin, with an emerging role for rifaximin. Oral rifaximin, a semisynthetic rifamycin derivative, is an effective and well-tolerated antibacterial for the management of adults with non-invasive traveller's diarrhea. Rifaximin was significantly more effective than placebo and no less effective than ciprofloxacin in reducing the duration of diarrhea. While rifaximin is effective in patients with E. coli-predominant traveller's diarrhea, it appears ineffective in patients infected with inflammatory or invasive enteropathogens.[81]
Prevention[edit]
ETEC is the type of E. coli that most vaccine development efforts are focused on. Antibodies against the LT and major CFs of ETEC provide protection against LT-producing, ETEC-expressing homologous CFs. Oral inactivated vaccines consisting of toxin antigen and whole cells, i.e. the licensed recombinant cholera B subunit (rCTB)-WC cholera vaccine Dukoral, have been developed. There are currently no licensed vaccines for ETEC, though several are in various stages of development.[82] In different trials, the rCTB-WC cholera vaccine provided high (85–100%) short-term protection. An oral ETEC vaccine candidate consisting of rCTB and formalin inactivated E. coli bacteria expressing major CFs has been shown in clinical trials to be safe, immunogenic, and effective against severe diarrhoea in American travelers but not against ETEC diarrhoea in young children in Egypt. A modified ETEC vaccine consisting of recombinant E. coli strains over-expressing the major CFs and a more LT-like hybrid toxoid called LCTBA, are undergoing clinical testing.[83][84]
Other proven prevention methods for E. coli transmission include handwashing and improved sanitation and drinking water, as transmission occurs through fecal contamination of food and water supplies. Additionally, thoroughly cooking meat and avoiding consumption of raw, unpasteurized beverages, such as juices and milk are other proven methods for preventing E.coli. Lastly, avoid cross-contamination of utensils and work spaces when preparing food.[85]
Model organism in life science research[edit]
Because of its long history of laboratory culture and ease of manipulation, E. coli plays an important role in modern biological engineering and industrial microbiology.[86] The work of Stanley Norman Cohen and Herbert Boyer in E. coli, using plasmids and restriction enzymes to create recombinant DNA, became a foundation of biotechnology.[87]
E. coli is a very versatile host for the production of heterologousproteins,[88] and various protein expression systems have been developed which allow the production of recombinant proteins in E. coli. Researchers can introduce genes into the microbes using plasmids which permit high level expression of protein, and such protein may be mass-produced in industrial fermentation processes. One of the first useful applications of recombinant DNA technology was the manipulation of E. coli to produce human insulin.[89]
Many proteins previously thought difficult or impossible to be expressed in E. coli in folded form have been successfully expressed in E. coli. For example, proteins with multiple disulphide bonds may be produced in the periplasmic space or in the cytoplasm of mutants rendered sufficiently oxidizing to allow disulphide-bonds to form,[90] while proteins requiring post-translational modification such as glycosylation for stability or function have been expressed using the N-linked glycosylation system of Campylobacter jejuni engineered into E. coli.[91][92][93]
Modified E. coli cells have been used in vaccine development, bioremediation, production of biofuels,[94] lighting, and production of immobilised enzymes.[88][95]
Strain K-12 is a mutant form of E. coli that over-expresses the enzyme Alkaline Phosphatase (ALP).[96] The mutation arises due to a defect in the gene that constantly codes for the enzyme. A gene that is producing a product without any inhibition is said to have constitutive activity. This particular mutant form is used to isolate and purify the aforementioned enzyme.[96]
Strain OP50 of Escherichia coli is used for maintenance of Caenorhabditis elegans cultures.
Strain JM109 is a mutant form of E. coli that is recA and endA deficient. The strain can be utilized for blue/white screening when the cells carry the fertility factor episome[97] Lack of recA decreases the possibility of unwanted restriction of the DNA of interest and lack of endA inhibit plasmid DNA decomposition. Thus, JM109 is useful for cloning and expression systems.
Model organism[edit]
E. coli is frequently used as a model organism in microbiology studies. Cultivated strains (e.g. E. coli K12) are well-adapted to the laboratory environment, and, unlike wild-type strains, have lost their ability to thrive in the intestine. Many laboratory strains lose their ability to form biofilms.[98][99] These features protect wild-type strains from antibodies and other chemical attacks, but require a large expenditure of energy and material resources. E. coli is often used as a representative microorganism in the research of novel water treatment and sterilisation methods, including photocatalysis. By standard plate count methods, following sequential dilutions, and growth on agar gel plates, the concentration of viable organisms or CFUs (Colony Forming Units), in a known volume of treated water can be evaluated, allowing the comparative assessment of materials performance. [100]
In 1946, Joshua Lederberg and Edward Tatum first described the phenomenon known as bacterial conjugation using E. coli as a model bacterium,[101] and it remains the primary model to study conjugation.[102]E. coli was an integral part of the first experiments to understand phage genetics,[103] and early researchers, such as Seymour Benzer, used E. coli and phage T4 to understand the topography of gene structure.[104] Prior to Benzer's research, it was not known whether the gene was a linear structure, or if it had a branching pattern.[105]
E. coli was one of the first organisms to have its genome sequenced; the complete genome of E. coli K12 was published by Science in 1997.[51]
By evaluating the possible combination of nanotechnologies with landscape ecology, complex habitat landscapes can be generated with details at the nanoscale.[106] On such synthetic ecosystems, evolutionary experiments with E. coli have been performed to study the spatial biophysics of adaptation in an island biogeography on-chip.
Studies are also being performed attempting to program E. coli to solve complicated mathematics problems, such as the Hamiltonian path problem.[107]
History[edit]
In 1885, the German-Austrian pediatrician Theodor Escherich discovered this organism in the feces of healthy individuals. He called it Bacterium coli commune because it is found in the colon. Early classifications of prokaryotes placed these in a handful of genera based on their shape and motility (at that time Ernst Haeckel's classification of bacteria in the kingdom Monera was in place).[84][108][109]
Bacterium coli was the type species of the now invalid genus Bacterium when it was revealed that the former type species ('Bacterium triloculare') was missing.[110] Following a revision of Bacterium, it was reclassified as Bacillus coli by Migula in 1895[111] and later reclassified in the newly created genus Escherichia, named after its original discoverer.[112]
Bacterium coli has since been used for biological lab experiment research. Infection can lead to hemolytic uremic syndrome (HUS), characterized by hemolytic anemia, thrombocytopenia, and renal injury.[113]
In 1996 the world's worst outbreak of E. coli food poisoning occurred in Wishaw, Scotland, killing 20 people.[114]
See also[edit]References[edit]
Databases and external links[edit]
Retrieved from 'https://en.wikipedia.org/w/index.php?title=Escherichia_coli&oldid=901904312'
Comments are closed.
|
Details
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
- Blog
- Ir Conversie Spiegelreflex Camera
- Venmar Ea 1500 Filter
- Son Rui Nodame
- Spss Version 25
- Maxwell Sketchup Torrent
- Tiny People Stomp
- Wondershare Keygen Mac
- Ivc Class Schedule
- Linux Ftdi Vcp
- Johnson Plastic Group, Sa De Cv
- Adobe Premiere Pro Cc 2018 Free Download Full Version With Crack
- Technic Launcher Pirata 64 Bits
- Download Pizza Frenzy For Mac
- Martin Backpacker Guitar Sound Hole Curve?
- Lineage 2 Private Server Bot Allowed
- New Camry Or Used Lexus
- The wolf among us season 2 writers
- Learn paint shop pro 9
- My passport ultra for mac reviews
- The green inferno full movie online megavideo
- Ricoh 3300 printer driver
- Rufus os to usb
- Get microsoft student discount
- Sims 4 rigs list
- Star wars the old republic wiki
- Firefox free download
- How to make a bootable usb mac os sierra
- Kitchen design software free home depot
 RSS Feed
RSS Feed
